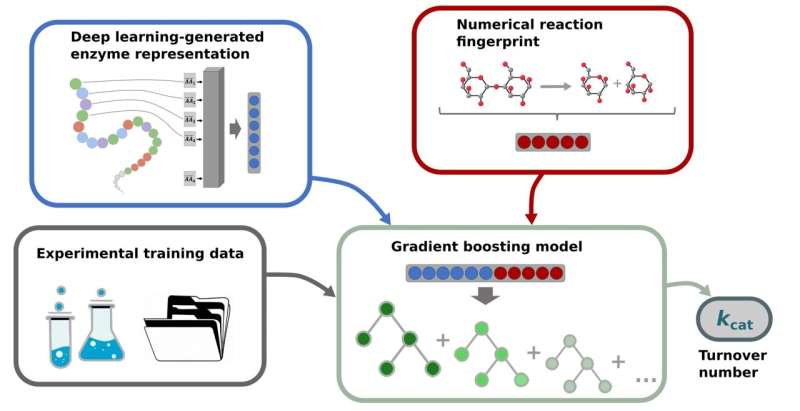
Schematic diagram of the prediction process for the turnover number of enzymatic reactions: Enzymes are sequences of amino acids; these sequences are converted into numerical vectors, depicted as gray squares, which are then transformed into a vector by a deep learning model (top left). Information about catalysed reactions is also generated as numerical vectors (top right). The experimentally determined turnover numbers (bottom left) are used to train a gradient boosting model to predict the kcat turnover number (bottom right). Gradient boosting models are an ensemble of multiple decision trees, depicted in different green tones. Source: HHU/Alexander Kroll
Enzymes play an important role in cellular metabolic processes. To make a quantitative assessment of these processes, researchers need to know the so-called “turnover number” (in short: kdespising) of enzymes. In the journal Communication in Naturea group of bioinformaticians from Heinrich Heine University Düsseldorf (HHU) has now described a tool for predicting this parameter for various enzymes using AI methods.
Enzymes are important biocatalysts in all living cells. They are usually large proteins, which bind to small molecules—called substrates—and then convert them into other molecules, the “products.”
Without enzymes, the reaction that converts substrates into products does not occur, or it occurs only at a very low rate. Most organisms have thousands of different enzymes. Enzymes have many applications in a wide range of biotechnological processes and in everyday life—from proofing bread dough to detergents.
The maximum speed at which a specific enzyme can convert its substrates into products is determined by the so-called turnover number kdespising. It is an important parameter for quantitative research of enzyme activities and plays an important role in understanding cellular metabolism.
However, determining k is time-consuming and expensivedespising turnover numbers in experiments, so they are unknown to most of the reactions. The Computational Cell Biology research group at HHU led by Professor Dr. Martin Lercher has now developed a new tool called TurNuP to predict kdespising turnover numbers of enzymes using AI methods.
In training akdespising prediction model, information about enzymes and catalyzed reactions are converted into numerical vectors using deep learning models. These numerical vectors serve as input for a machine learning model—a so-called gradient boosting model—that predicts kdespising turnover numbers.
Lead author Alexander Kroll said, “TurNuP outperforms previous models and can even be used successfully for enzymes with little similarity to those in the training dataset.” Previous models did not make any meaningful predictions unless at least 40% of the enzyme sequence was identical to at least one enzyme in the training set. In contrast, TurNuP can now make meaningful predictions for enzymes with a maximum sequence identity of 0 to 40%.
Professor Lercher added, “In our study, we show that the predictions made by TurNuP can be used to predict the concentrations of enzymes in living cells more accurately than is currently the case.”
To make the prediction model easily accessible to as many users as possible, the HHU team has developed a user-friendly web server, which can be used by other researchers to predict kdespising turnover rate of enzymes.
More information:
Alexander Kroll et al, Turnover number predictions for kinetically uncharacterized enzymes using machine and deep learning, Communication in Nature (2023). DOI: 10.1038/s41467-023-39840-4
Provided by Heinrich Heine University Düsseldorf
Citation: AI tool predicts work rate of enzymes (2023, July 24) retrieved on 25 July 2023 from https://phys.org/news/2023-07-ai-tool-enzymes.html
This document is subject to copyright. Except for any fair dealing for the purpose of private study or research, no part may be reproduced without written permission. Content is provided for informational purposes only.
